Our keywords: Virology, neurobiology, cell biology, synaptic plasticity, intracellular trafficking, neurodegeneration, live cell imaging, machine learning, antivirals, secretion, extracellular vesicles, … Viruses we particularly like these days: Coronaviruses, Flaviviruses, Bunyaviruses, Hepatitis B virus, HIV, … The team members are working together on interdisciplinary fields in order to tackle major questions associated to virology, cell biology and neurobiology. Most precisely, we are oriented around three major axis :
- Membrane dynamics of viral and cellular components
- Impact of viruses on neural functions
- Technological development & translational research
All three are developed a bit further down. Stay with us ! 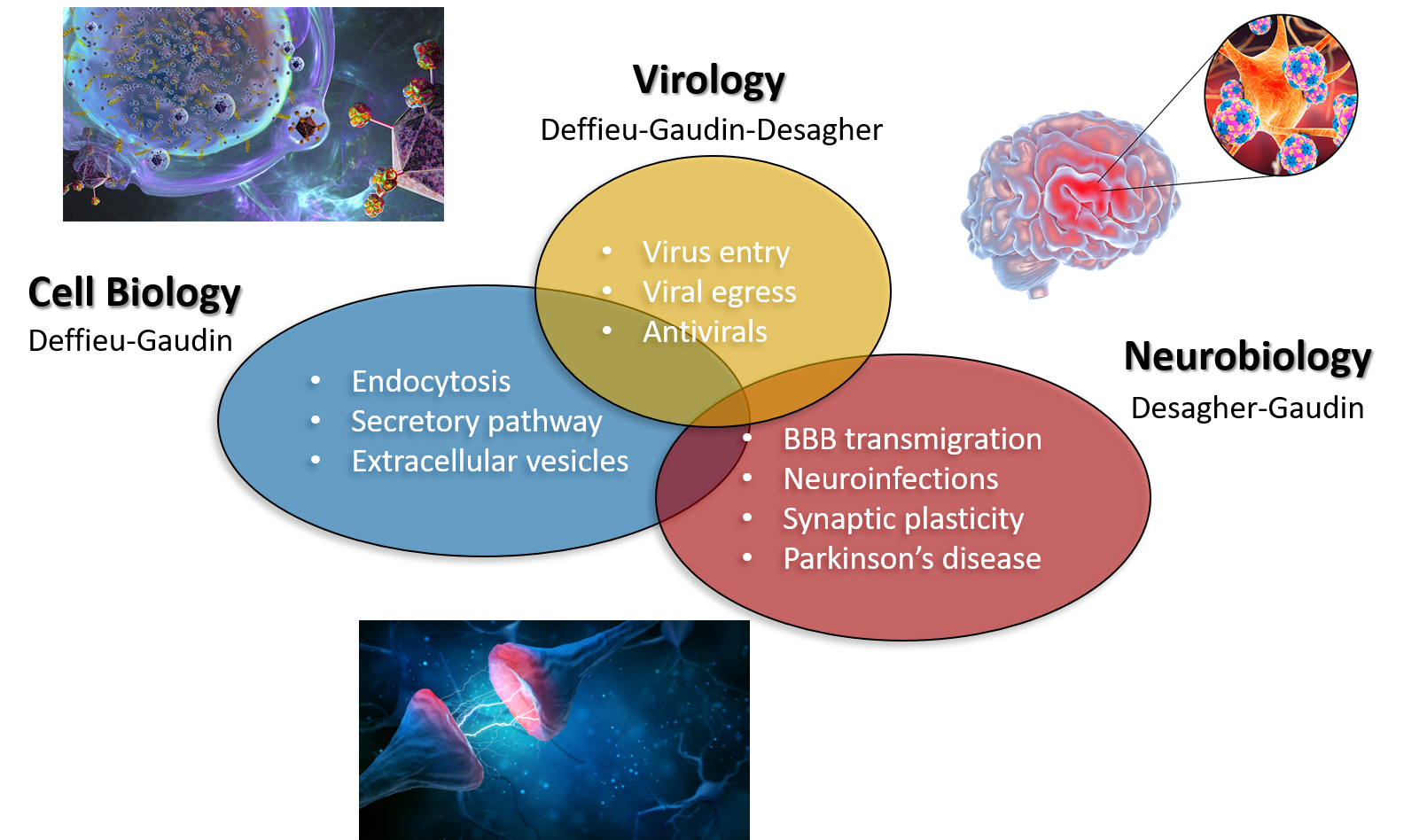
Membrane dynamics of viral and cellular components
Hepatitis B virus
Coord: Deffieu. Involved lab members: Gaudin, Bohbot, Arroyo.
We are studying Hepatitis B virus (HBV) entry and egress from hepatocytes. Our work aims to characterize every step of HBV trafficking. At the entry step, we are deciphering the molecular mechanisms involved in HBV attachment to its plasma membrane receptors, and tracking its kinetics of escape from the endosomes. At the egress step, we are monitoring the dynamics of particle assembly and secretion and characterizing the host proteins involved in this process. These different mechanisms are monitored by tracking HBV viral dynamic at the scale of the single particle using fluorescent microscopy.
Cellular pathways associated with secretion (biosynthetic pathway and extracellular vesicles)
Coord: Deffieu. Involved lab members: Gaudin.
We recently discovered a new secretory pathway involving the small Rab7A-GTPase (Deffieu et al, Sci Adv, 2022). To determine the role of the Rab7A secretory pathway, we are investigating its molecular partners and the receptors transiting through this route. We aim on characterizing its physiological function in non-polarized and polarized cells. Our goal is to develop a specific methodology to investigate new transport mechanisms involved in the biosynthetic secretory pathway and for extracellular vesicle (EV) release.
Coronavirus entry
Coord: Gaudin. Involved lab members: Lebrun, Cogrossi, Gorda.
We are deciphering how SARS-CoV-2 are breaking into target cells through the development of new molecular tools for the visualization of virus attachment internalization and fusion with cellular membranes. This methodology allows us to decipher the role of host factors in the early stages of infection.
ER remodelling upon infection
Coord: Gaudin. Involved lab members: Deffieu, Bouget, Lebrun, Cogrossi.
Replication of flaviviruses and coronaviruses occurs in the endoplasmic reticulum (ER), which leads to dramatic subcellular remodeling. Our lab investigates how viruses modulate ER functions and underlying mechanisms in cell lines and neurons. Moreover, we have designed a series of proprietary small molecules targeting the ER, which exhibit potent broad-spectrum antiviral properties. Using advanced techniques in microscopy and mass spectrometry, we are characterizing the mode of action and target of these compounds.
Impact of viruses on neural functions
Parkinson’s disease
Coord: Desagher. Involved lab members: Gaudin, Kumarasinghe, Gaudillat.
Epidemiological studies have established a link between viral infections and neurodegenerative diseases, notably between influenza and Parkinson’s disease. In order to determine whether infection of neurons with influenza virus is a triggering event in the pathogenesis of Parkinson’s disease, we study the impact of this virus on alpha-synuclein, a protein whose accumulation and aggregation plays a major role in this disease.
Synaptic plasticity
Coord: Gaudin. Involved lab members: Desagher, Bouget, Almeida, Béchard, Gorda, Arroyo, Saini.
Viruses significantly impact neurological functions, causing life-threatening encephalitis as well as more subtle neurocognitive symptoms, as we recently reviewed (Bouget et al, Trends Neurosci, 2025). Our lab investigates how exposure to viruses can lead to synaptic impairment using a combination of approaches, including high-resolution imaging, electrophysiology, mass spectrometry. To better represent the complexity of the brain architecture, we are employing brain organoids and organotypic culture of human post-mortem adult brain explants (OPAB), which exhibit relevant circuitry and plasticity.
Perturbation of neural functions upon HIV exposure
Coord: Gaudin. Involved lab members: Lutz, Béchard, Brychka, Ouedraogo.
Current antivirals are very potent to inhibits HIV infection, but no drug exists to completely clear the virus from infected individuals. The brain is a major reservoir for the virus’ persistence, and HIV-associated neurocognitive disorders (HAND), have been extensively described. Through the screening of infected brain organoid by confocal imaging coupled to the development of a novel self-supervised machine learning framework (SSL), we are trying to predict and classify HIV-specific perturbations of neuron and astrocyte morphology and mitigation strategies. Combined to scRNAseq, we aim to determine main pathways involved in HAND and ways to counteract it, using symptom-oriented readouts.
Crossing the Blood-Brain Barrier
Coord: Gaudin. Involved lab members : Arroyo, Béchard, Krahel, Saini.
Viruses can break into the brain through different ways, including the Trojan horse strategy. Our lab is studying how viruses hijack host machineries and cells to cross the endothelial cells composing the blood-brain barrier (BBB). Furthermore, even in the absence of brain invasion, viruses are able to perturb significantly the neurovascular unit (NVU), leading to indirect neurological perturbations. Using transwells and microfluidic approaches, our goal is to provide mechanistic insights as to how viruses perturbs the endothelium-brain crosstalk.
Technological development & translational research
3D neural models
Coord:Gaudin. Involved lab members: Desagher, Bouget, Gorda, Almeida.
Our lab is using human induced pluripotent stem cells (hiPSCs) to grow brain organoids, small balls of neurons, astrocytes, and neural progenitor cells, exhibiting basic fetal brain architecture. Over the past years, we also developed and standardized the organotypic culture of post-mortem adult brain explants (OPAB), capable to remain alive in culture for months (Partiot, Gorda, et al, EMBO Mol Med, 2024). These 3D neural models are used to evaluate the molecular mechanisms by which viruses subvert host machineries and cause neural dysfunctions.
Antivirals
Coord: Gaudin. Involved lab members: Deffieu, Cogrossi, Lebrun, Gorda, Lutz, Béchard.
During our mechanistic investigations, our lab aims to convert some of the findings into potential novel therapeutic strategies. In particular, we have designed a series of proprietary small molecules targeting the ER, which exhibit potent broad-spectrum antiviral properties. Using advanced techniques in microscopy and mass spectrometry, we are characterizing the mode of action and target of these compounds. Moreover, we are developing novel AI-based pipelines to predict drug potency and identify novel symptom-oriented antivirals. Finally, we are actively collaborating with industries to evaluate the antiviral potency of natural products.
Biophysics of virus-containing saliva
Coord:Gaudin. Involved lab members: Cogrossi, Ouedraogo, Gorda, Lebrun.
Our lab aims to understand how saliva composition affects SARS-CoV-2 infectiveness. Indeed, it appears that some people are highly contagious (superspreaders), while others are not, an observation that is not explained by the viral load in saliva. Using clinical saliva samples from infected and non-infected donors, we aim to identify the determinants of SARS-CoV-2 infectiveness.
Bioinformatics and Artificial Intelligence
Coord: Gaudin. Involved lab members: Lutz, Ouedraogo.
Our lab has developed a user-friendly software to unbiasedly analyze electrophysiological recordings from neurons, organoids, and brain explants in-a-dish, and electroencephalograms (EEG) from patients. It is based on a simple random forest classifier (RFC) framework that is sufficient for the type of data recorded. We are also implementing more sophisticated self-supervised learning (SSL) algorithms for the analyses of microscopy images, through the generation of our own foundation model. Moreover, our lab has invested significant efforts into the analysis of single cell datasets, including scRNAseq, and single cell Proteomics (scP). While scRNAseq analyses are already quite well established, scP suffers from the lack of validated workflow, in particular for imputation, normalization, aggregation, and cell type labelling. We are thus committed to provide robust pipelines for in-depth analyses of scP.
Single cell (spatial) proteomics
Coord: Gaudin. Involved lab members: Almeida, Ouedraogo.
As part of a national consortium, our lab develops novel methodologies to perform advanced single cell spatial proteomics (scSP). In coordination with our collaborators, we are developing in parallel biological sample preparation, AI-based segmentation and isolation, and highly sensitive mass spectrometry to uncover how bunyaviruses interfere with the neural proteome.
Neurohybrid systems
Coord: Gaudin. Involved lab members: Béchard.
Our lab is collaborating on interdisciplinary projects aiming at reproducing the basic learning functions of the human brain in-a-dish. The ultimate goal is to explore how viruses can interfere with cognitive functions in animal-free experimentations.
Microfluidics
Coord: Gaudin. Involved lab members: Béchard.
We have recently implemented microfluidic devices in order to be able to inject and collect biological and chemical elements to brain explants while recording their electrophysiological activity. This approach is being extended to organoids and for the culture of BBB-like endothelial cells.
Molecular tools for live imaging
Coord: Deffieu. Involved lab members: Gaudin, Desagher, Lebrun, Gorda, Almeida, Bouget, Arroyo, Bohbot.
Our lab builds and/or implement the use of a large panel of reporter viruses (HBV, HCV, SARS-CoV-2, HIV, hCoV-229E, DENV, WNV, ZIKV, BUNV, …) fused to bright and photostable fluorescent proteins or luciferase to track virus entry, replication, and egress. In addition, we are using CRISPR-Cas9 knock-in strategies, the nanobody-ALFA complex, the APEX tag, and other original tags, to monitor protein trafficking and virus-host interactions in live cells. These tools are helping us uncover the underlying molecular mechanisms involved.
Plasmids from the team are available at:
Our team « Membrane Dynamics & Viruses » (MDV) aims at studying virus-host interactions at different spatiotemporal scales. The viruses studied in the lab allow us to better understand physiological molecular and cellular mechanisms, and vice-versa, the investigation of cellular processes provides valuable information regarding viral subversion mechanisms. Specifically, our projects are focusing on three complementary axes:
- The intracellular transport in/out: how do viruses (HBV, HCV, SARS-CoV-2, …) interact with their receptor at the cell surface of target cell? Conversely, how does the cell control neo-synthesized transmembrane viral receptor secretion and extracellular vesicles?
- How viruses (Zika and HIV) cross the blood-brain barrier? What is the impact of brain exposure to viruses (Flaviviruses, Coronaviruses, Bunyaviruses, HIV, …)?
- Development of novel antiviral molecules with broad spectrum. How to improve their potency, stability, solubility, bioavailability? What is the host protein targeted?
 In project I, we have identified that DNAseI is encapsidated into viral particles during HBV assembly (Hallez et al., Nature Microbiol, 2019) and imaged the association of Core-DNAseI in real time (Figure 1).
In project I, we have identified that DNAseI is encapsidated into viral particles during HBV assembly (Hallez et al., Nature Microbiol, 2019) and imaged the association of Core-DNAseI in real time (Figure 1).
Figure 1. 5D reconstruction of the spatiotemporal dynamics of HBV-Core (green) and DNAseI (red). As part of project II, we found that Zika virus and HIV enhance monocyte transmigration through the blood-brain barrier in vitro and in vivo. Our results reinforce the “Trojan Horse” hypothesis (Figure 2). 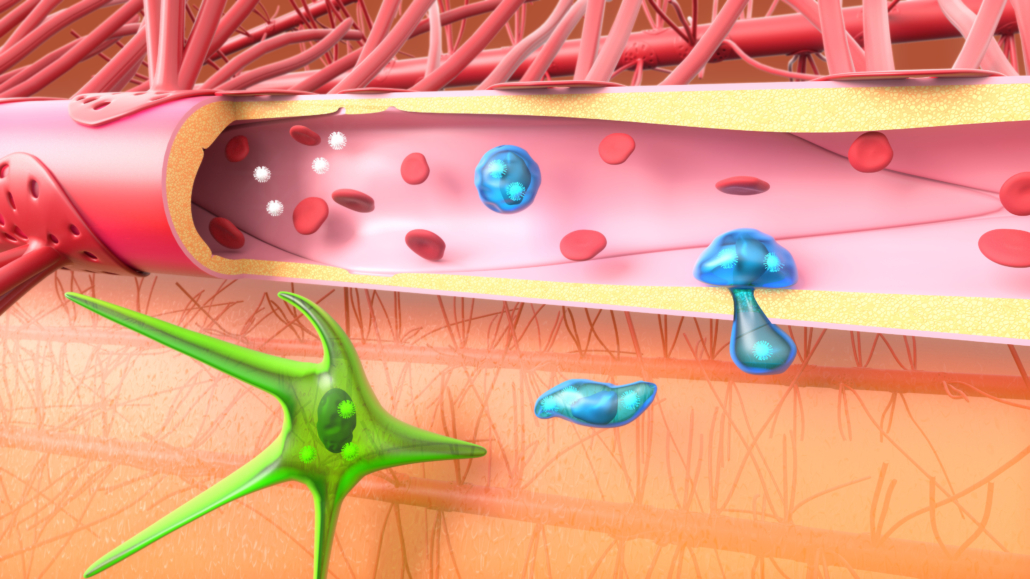 Figure 2. Viral particles in the bloodstream infect (or are captured by) blood cells, which will transmigrate through the endothelium, allowing the virus to cross this impermeable barrier. We are routinely employing techniques related to the study of HBV, HCV, HIV, and Zika viruses, including RT-qPCR, Western blot, flow cytometry, immunostaining, RNA interference, plaque assays, … We also developed advanced methods for the study of virus-host interactions such as CRISPR/Cas9 knock-in/out, RUSH, high resolution 3D confocal live cell imaging, mass spectrometry, xenotypic zebrafish embryo model system, human explants, cerebral orgaoids derived from stem cells, … We have implemented the culture of stem cell derived cerebral organoids in the lab (Figure 3).
Figure 2. Viral particles in the bloodstream infect (or are captured by) blood cells, which will transmigrate through the endothelium, allowing the virus to cross this impermeable barrier. We are routinely employing techniques related to the study of HBV, HCV, HIV, and Zika viruses, including RT-qPCR, Western blot, flow cytometry, immunostaining, RNA interference, plaque assays, … We also developed advanced methods for the study of virus-host interactions such as CRISPR/Cas9 knock-in/out, RUSH, high resolution 3D confocal live cell imaging, mass spectrometry, xenotypic zebrafish embryo model system, human explants, cerebral orgaoids derived from stem cells, … We have implemented the culture of stem cell derived cerebral organoids in the lab (Figure 3). 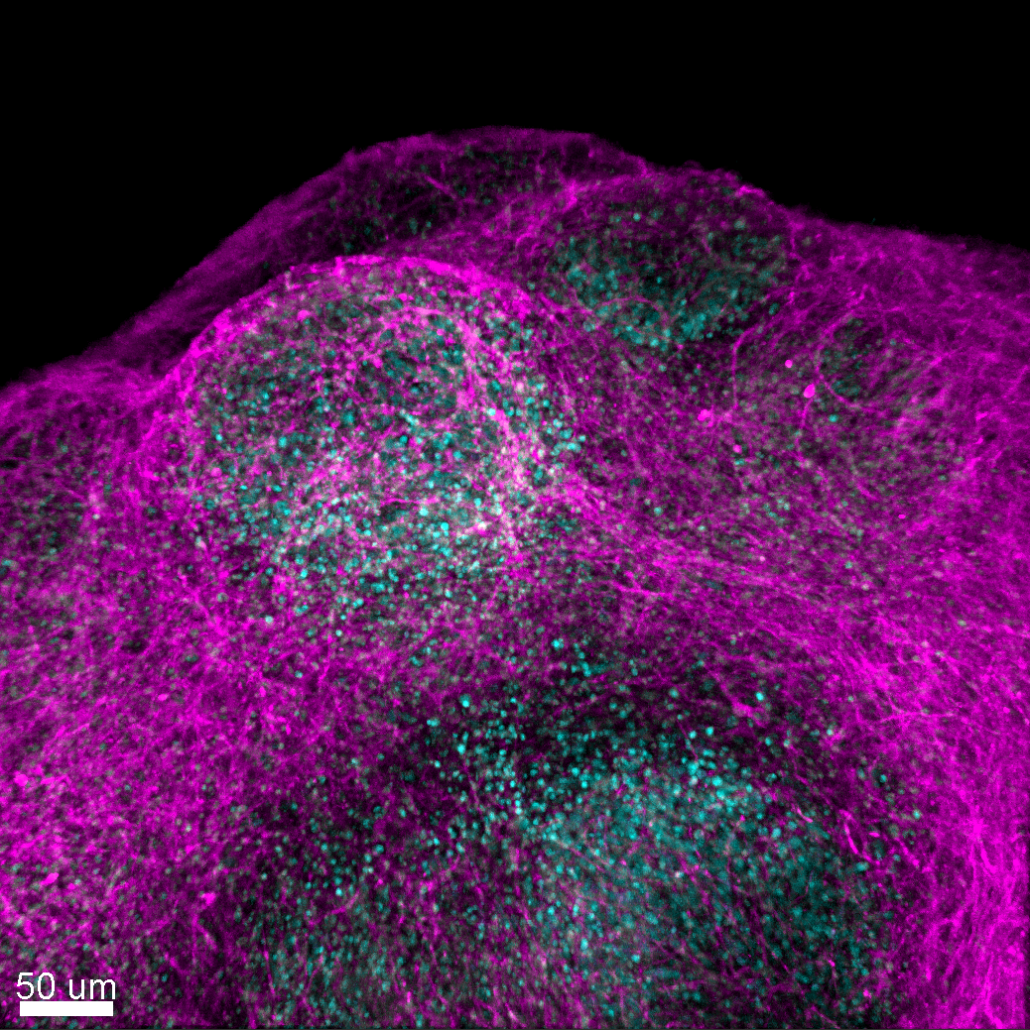
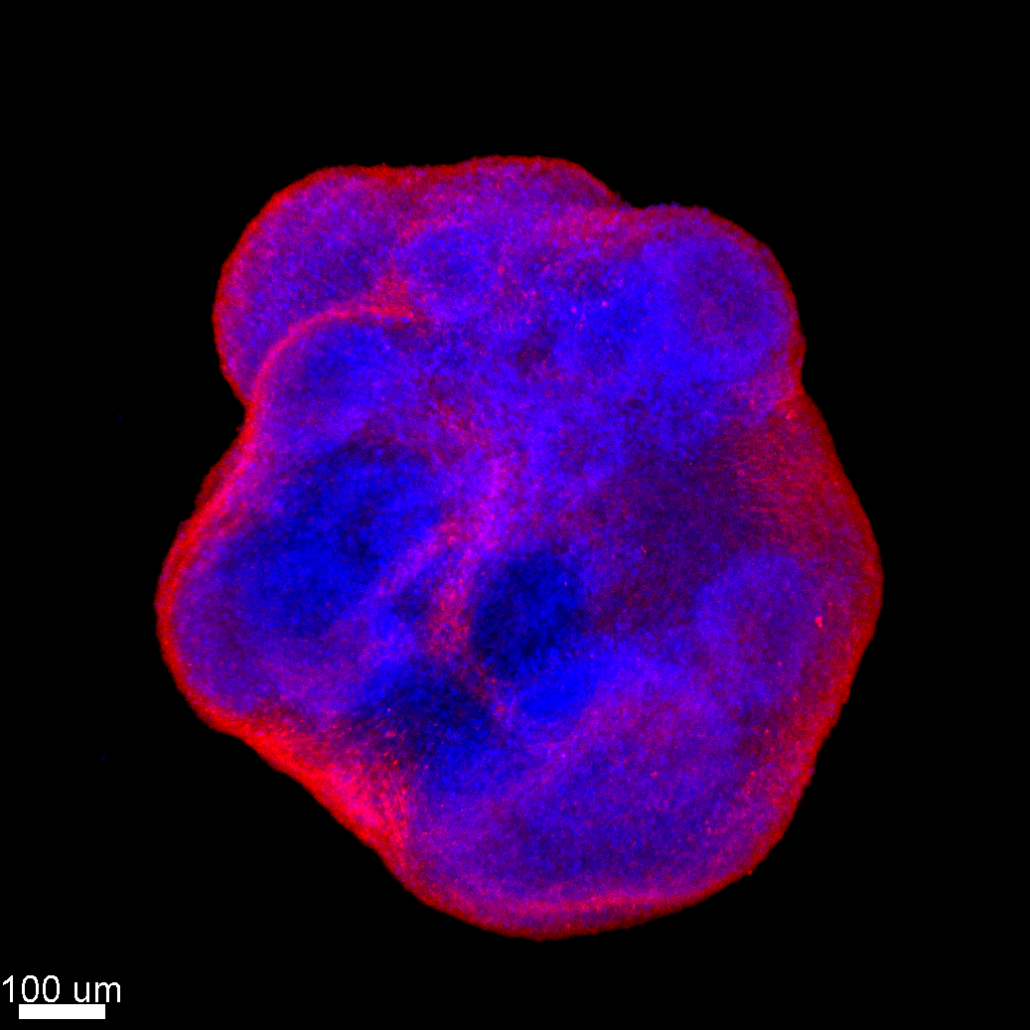 Figure 3. Confocal imaging of cerebral organoids
Figure 3. Confocal imaging of cerebral organoids
The philosophy of the lab is to go beyond technological limitations, not accepting “we can’t do it” as an answer, but rather “how are we going to achieve it?”.
The team is affiliated to the French society for cell biology (SBCF) and R.Gaudin is an elected member of the SBCF board council since 2016  R.Gaudin has been nominated as “full member” of the World Society for Virology (WSV) in 2019.
R.Gaudin has been nominated as “full member” of the World Society for Virology (WSV) in 2019. 
Current fundings
- PEPR MIE, coordinator R.Gaudin. Project ViroBrain.
- Recherche et Innovation à Risque (RI²), CNRS, coordinateur R.Gaudin. Project ProteoVir.
- ANRS, coordinator R.Gaudin. Project Barbie.
- ANRS PRFI, coordinator internationale avec le Brézil, coordinateur R.Gaudin. Project EviSyp.
- Human Frontiers Scientific Program (HFSP), coordinator R.Gaudin. Project DeViNe.
- ANR PRC, partner R.Gaudin. Project SCADRIV.
- ANR PRC, coordinator R.Gaudin. Project isiBrain.
- FRM, coordinator S.Desagher. Project Transcriptional regulation of alpha-synuclein in Parkinson’s disease.
- ANRS, coordinator M.Deffieu. Project Hepatitis B Virus.
Previous fundings
- ANR
- ANRS
- Sidaction
- CNRS-INSB
- SATT AxlR
- La Région Occitanie
- Labex NUMEV
- Atip-Avenir
- FRM
- EMBO LTF
- INSERM-région
- Chaire d’attractivité Idex (Université de Strasbourg)
- Prestige (Européen)
- Programme Stefanik (Ambassade Franco-Slovaque)
- Initiation of collaboration with China (franco-chinese ambassy)
- Atip-Avenir
At a glance
The MDV team studies what viruses do to cells… Or what cells do to viruses… Well, we study the intimate virus-cell relationship
Funding